import numpy as np
import pandas as pd
from sklearn import decomposition
from pymongo import MongoClient
from sklearn.pipeline import Pipeline
from sklearn.impute import KNNImputer
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.preprocessing import StandardScaler
from sklearn.compose import ColumnTransformer
from scipy import stats
from sklearn.preprocessing import PowerTransformer
import numpy as np
from sklearn.manifold import Isomap
import matplotlib.pyplot as plt
from sklearn.metrics import mean_absolute_percentage_error
import pandas as pd
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.preprocessing import PowerTransformer
# To use this experimental feature, we need to explicitly ask for it:
from sklearn.experimental import enable_iterative_imputer # noqa
from sklearn.datasets import fetch_california_housing
from sklearn.impute import SimpleImputer
from sklearn.impute import IterativeImputer
from sklearn.linear_model import BayesianRidge
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import ExtraTreesRegressor
from sklearn.neighbors import KNeighborsRegressor
from sklearn.pipeline import make_pipeline
from sklearn.model_selection import cross_val_score
import plotly.graph_objects as go
import plotly.tools as tls
from plotly.offline import plot, iplot, init_notebook_mode
from IPython.core.display import display, HTML
from plotly.subplots import make_subplots
from sklearn.model_selection import GridSearchCV
from sklearn.decomposition import TruncatedSVD
from xgboost.sklearn import XGBRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import cross_val_score
init_notebook_mode(connected = True)
config={'showLink': False, 'displayModeBar': False}
num_att = ['First Registration',
'Mileage',
'Power(hp)',
'Displacement']
X_num_att = ['First Registration',
'Mileage',
'Power(hp)',
'Displacement']
cat_att = ['Make', 'Model', 'Body', 'Fuel','Gearing Type']
class InitalCleaning(BaseEstimator, TransformerMixin):
def __init__(self):
pass
def fit(self, X, y=None):
return self
def transform(self, play_df, y = None):
play_df = play_df.drop(columns = ['ID', 'Loaded_in_DW', 'Model Code'])
#Adjust Column Types
play_df['Power(hp)'] = pd.to_numeric(play_df['Power(hp)'])
play_df['Displacement'] = pd.to_numeric(play_df['Displacement'])
play_df['Mileage'] = pd.to_numeric(play_df['Mileage'])
play_df['Price'] = pd.to_numeric(play_df['Price'])
#Drop rows with null values
play_df = play_df[~(play_df['Make'].isna() | play_df['Model'].isna())]
play_df = play_df[~(play_df['Displacement'].isna() & play_df['Power(hp)'].isna())]
play_df = play_df[~(play_df["Body"].isna())]
play_df = play_df[~(play_df["Fuel"].isna())]
play_df = play_df[~(play_df["Price"].isna())]
#Drop fake ads and update column values
play_df = play_df[~(play_df['Displacement'] < 900)]
play_df = play_df[~(play_df['Displacement'] > 70000)]
play_df = play_df[~((play_df['Power(hp)']>500) & \
(~((play_df['Make'] == 'Audi') | (play_df['Make'] == 'BMW') | (play_df['Make'] == 'Mercedes-Benz'))))]
play_df = play_df[~((play_df['Power(hp)']<30) & (~(play_df['Fuel'] == 'Electricity')))]
play_df = play_df[~(play_df['Price']>300000)]
play_df["Fuel"] = play_df["Fuel"].str.split("/").str[0]
play_df['Gearing Type'] = play_df['Gearing Type'].replace({np.nan : 'Manual'})
return play_df
class AdjustEquip(BaseEstimator, TransformerMixin):
def __init__(self):
pass
def fit(self, X, y=None):
return self
def transform(self, play_df, y = None):
#Keep only 0 and 1 values
equipment = play_df.iloc[:,9:79]
equipment = equipment.replace({np.nan: 0})
equipment = equipment.replace({'1': 1})
a = set(equipment['Warranty'])
a.remove(0)
a.remove(1)
equipment = equipment.replace(list(a) , 1)
#Convert each column's type to int
for col in equipment.columns:
equipment[col] = equipment[col].astype(int)
play_df.iloc[:,9:79] = equipment
return play_df
class CategoricalEncoder(BaseEstimator, TransformerMixin):
def __init__(self):
pass
def fit(self, X, y=None):
return self
def transform(self, play_df, y = None):
play_df = pd.get_dummies(data = play_df,
columns = ['Make', 'Model', 'Body', 'Fuel','Gearing Type'],
dummy_na = False)
return play_df
class RegistrationTransformer(BaseEstimator, TransformerMixin):
def __init__(self, tip = 0):
self.tip = tip
def fit(self, X, y=None):
return self
def transform(self, play_df, y = None):
#Reset index is used to avoid null values when merging
play_df.reset_index(drop=True, inplace=True)
a = date_magic(play_df['First Registration'])
a.reset_index(drop = True, inplace = True)
play_df['First Registration'] = a
if self.tip == 0:
play_df = play_df[(play_df['Make']=='BMW') & (play_df['First Registration'] > 2005) & ( (play_df['Model'].str.startswith('3')) | (play_df['Model'].str.startswith('1')) ) ]
elif self.tip == 1:
play_df = play_df[play_df.isnull().any(axis=1)]
return play_df
class IQROutlierRemoval_new(BaseEstimator, TransformerMixin):
def __init__(self, num_att):
self.num_att = num_att
def fit(self, X, y = None):
return self
def transform(self, play_df, y = None):
# Q1 = play_df.quantile(0.25)
# Q3 = play_df.quantile(0.75)
# IQR = Q3- Q1
# play_df = play_df[~((play_df < (Q1 - 1.5 * IQR)) |(play_df > (Q3 + 1.5 * IQR))).any(axis=1)]
return pd.DataFrame(play_df, columns = num_att)
class IQROutlierRemoval(BaseEstimator, TransformerMixin):
def __init__(self, num_att):
self.num_att = num_att
def fit(self, X, y = None):
return self
def transform(self, play_df, y = None):
bmw_num = play_df[self.num_att]
bmw_cat = play_df.drop(self.num_att, axis = 1)
Q1 = bmw_num.quantile(0.25)
Q3 = bmw_num.quantile(0.75)
IQR = Q3- Q1
bmw_num = bmw_num[~((bmw_num < (Q1 - 1.5 * IQR)) |(bmw_num > (Q3 + 1.5 * IQR))).any(axis=1)]
return pd.DataFrame(bmw_num, columns = num_att)
class ZScoreOutlierRemoval(BaseEstimator, TransformerMixin):
def __init__(self, num_att):
self.num_att = num_att
def fit(self, X, y = None):
return self
def transform(self, play_df, y = None):
z = np.abs(stats.zscore(play_df, nan_policy='omit'))
return play_df[(z < 3).all(axis=1)]
class StandardScalerIndices(BaseEstimator, TransformerMixin):
def __init__(self):
self.scaler = StandardScaler()
def fit(self, X, y=None):
self.scaler.fit(X)
return self
def transform(self, play_df, y = None):
return pd.DataFrame(self.scaler.transform(play_df),columns = play_df.columns, index = play_df.index)
class PowerTransformerIndices(BaseEstimator, TransformerMixin):
def __init__(self, method):
self.method = method
self.transformer = PowerTransformer(method = self.method)
def fit(self, X, y=None):
self.transformer.fit(X)
return self
def transform(self, play_df, y = None):
return pd.DataFrame(self.transformer.transform(play_df),columns = play_df.columns, index = play_df.index)
class RemovePrice(BaseEstimator, TransformerMixin):
def __init__(self):
pass
def fit(self, X, y = None):
return self
def transform(self, play_df, y = None):
return play_df.drop('Price', axis = 1)
class IterativeImputerIndices(BaseEstimator, TransformerMixin):
def __init__(self, estimator):
self.imputer = IterativeImputer(estimator = estimator)
def fit(self, X, y=None):
self.imputer.fit(X)
return self
def transform(self, play_df, y = None):
return pd.DataFrame(self.imputer.transform(play_df),columns = play_df.columns, index = play_df.index)
class JoinTransformer(BaseEstimator, TransformerMixin):
def __init__(self, BMW_df, new_att):
self.BMW_df = BMW_df
self.new_att = new_att
def fit(self,X,y = None):
return self
def transform(self, X, y = None):
new = self.BMW_df.drop(self.new_att, axis = 1)
return X.join(new)
def date_magic(d):
months_dict = {'Jan': 1,'Feb':2,'Mar':3,'Apr':4 ,'May':5 ,'Jun':6,'Jul':7,
'Aug':8,'Sep':9,'Oct':10 ,'Nov':11,'Dec':12}
year = []
month = []
num_year = []
for el in d:
el = str(el)
split = el.split('-')
if (len(split)>1):
month.append(months_dict[split[0]])
if(int(split[1])>20):
year.append(int('19' + split[1]))
else:
year.append(int('20' + split[1]))
continue
split = el.split('/')
if(len(split)>1):
month.append(int(split[0]))
year.append(int(split[1]))
continue
month.append(1)
year.append(int(el))
month = pd.Series(month)
year = pd.Series(year)
num_year = year + month/12
return pd.Series(num_year)
def readData():
client = MongoClient('mongodb+srv://<User>:<Pass>@dwprojectcluster.lpqbf.mongodb.net/cars_database?retryWrites=true&w=majority')
df_cars = pd.DataFrame(list(client.cars_database.cars.find({})))
df_cars.drop('_id', axis = 1, inplace = True)
df_cars = df_cars[df_cars['Loaded_in_DW'].eq(False)]
df_cars.drop_duplicates(subset=['ID'], inplace = True)
return df_cars
recnik = {}
recnik['Method'] = []
recnik['Mean Percentage Error'] = []
recnik['Standard Deviation'] = []
def randomforestCV( max_features, n_estimators, X, y, message, scoring = 'neg_mean_absolute_percentage_error', recnik = recnik):
rfr = RandomForestRegressor(max_features = max_features, n_estimators = n_estimators, random_state = 2)
rfr.fit(X,y)
rfr_scores = cross_val_score(rfr, X, y, scoring = scoring, cv = 5, n_jobs = -1)
print("Scores:", -rfr_scores)
print("Mean:", -rfr_scores.mean())
print("Standard deviation:", rfr_scores.std())
recnik['Method'].append(message)
recnik['Mean Percentage Error'].append(-rfr_scores.mean())
recnik['Standard Deviation'].append(rfr_scores.std())
def gridSearch(param_grid, model, X, y):
grid_search = GridSearchCV(model, param_grid, cv=5,
scoring='neg_mean_absolute_percentage_error',
return_train_score=True,
verbose = 10, n_jobs = -1)
grid_search.fit(X, y)
cvres = grid_search.cv_results_
for mean_score, params in zip(cvres["mean_test_score"], cvres["params"]):
print(-mean_score, params)
return grid_search
1. Reading the data and inital data preprocessing
The data is being read from a MongoDB database which is being updated daily with new entires from the AutoScout24 website. First we are removing the entires that are missing values for some fields that are mandatory and can not be imputed. Next some of the outliers are removed manually by checking for values that don't make any sense.
The date is being transformed in a continious, decimal value ranging from 1990 to 2020, where the decimal represents the month of the year.
Given the limited computational power, we will reduce the dataset only to BMW cars first registered no earlier than 2005. The other 2 pipelines are used for visualization purposes and selecting the best imputation method respectively.
df_cars = readData()
type0_pipeline = Pipeline([
('initial', InitalCleaning()),
('equipment', AdjustEquip()),
('date', RegistrationTransformer(tip = 0))
])
type1_pipeline = Pipeline([
('initial', InitalCleaning()),
('equipment', AdjustEquip()),
('date', RegistrationTransformer(tip = 1)),
('encoder', CategoricalEncoder())
])
no_category = Pipeline([
('initial', InitalCleaning()),
('equipment', AdjustEquip()),
('date', RegistrationTransformer(tip = 0)),
])
BMW_df = type0_pipeline.fit_transform(df_cars)
nan_df = type1_pipeline.fit_transform(df_cars)
viz_df = no_category.fit_transform(df_cars)
2. Spliting the dataset to test and train set
Since we will be using k-fold cross validation when evaluating the models, this part of spliting the data to train and test set is not necessary(hence the value of 0.01 for test size). However this part is left as an example for stratified sampling, where the model of the car is taken into account when spliting the dataset.
BMW_df = BMW_df.reset_index()
BMW_df = BMW_df.drop('index', axis = 1)
split = StratifiedShuffleSplit(n_splits=1, test_size=0.01, random_state=42)
for train_index, test_index in split.split(BMW_df, BMW_df["Model"]):
strat_train_set = BMW_df.loc[train_index]
strat_test_set = BMW_df.loc[test_index]
y_train = strat_train_set['Price']
x_train = strat_train_set.drop('Price', axis = 1)
y_test = strat_test_set['Price']
x_test = strat_test_set.drop('Price', axis = 1)
3. Visualizing the data - Click HERE for the output
From the results we can get a better picture about the nature of our dataset. The most interesting and valuable part here are the correlations between different features. As expected we can see that there is strong negative linear correlation between the mileage of the car and it's first registration date. Said in simple terms, the older the car, the more miles it is likely to have passed. This rule is even more obvious when seeing the nonlinear correlation between these two variables, obtained from the Spearman's correlation matrix. Also the mileage affects the price negatively. The features that that have the biggest positive impact on the price are the power in hp and the age of the car. Suprsingly the displacement of the car is weakly correlated to the price. An interesting correlation is found between the model of the car and it's horsepower.
import pandas_profiling as pp
from plotly.offline import plot, iplot, init_notebook_mode
from IPython.core.display import display, HTML
init_notebook_mode(connected = True)
config={'showLink': False, 'displayModeBar': False}
pp.ProfileReport(viz_df.iloc[:,np.r_[0:10]]).to_file('aa.html')
N_SPLITS = 5
rng = np.random.RandomState(0)
X = nan_df.drop(columns = ['Price'])
y = nan_df['Price']
X = X.to_numpy()
y = y.to_numpy()
score_simple_imputer = pd.DataFrame()
br_estimator = BayesianRidge()
for strategy in ('mean', 'median'):
estimator = make_pipeline(
SimpleImputer(missing_values=np.nan, strategy=strategy),
br_estimator
)
score_simple_imputer[strategy] = cross_val_score(
estimator, X, y, scoring='neg_mean_squared_error',
cv=N_SPLITS
)
score_simple_imputer
X = X[::5]
y = y[::5]
estimators = [
DecisionTreeRegressor(max_features='sqrt', random_state=0),
ExtraTreesRegressor(n_estimators=10, random_state=0),
KNeighborsRegressor(n_neighbors=15),
BayesianRidge()
]
score_iterative_imputer = pd.DataFrame()
for impute_estimator in estimators:
estimator = make_pipeline(
IterativeImputer(random_state=0, estimator=impute_estimator),
br_estimator
)
score_iterative_imputer[impute_estimator.__class__.__name__] = \
cross_val_score(
estimator, X, y, scoring='neg_mean_squared_error', verbose = 2,
cv=N_SPLITS
)
score_iterative_imputer
;
scores = pd.concat(
[score_simple_imputer, score_iterative_imputer],
keys=['SimpleImputer', 'IterativeImputer'], axis=1
)
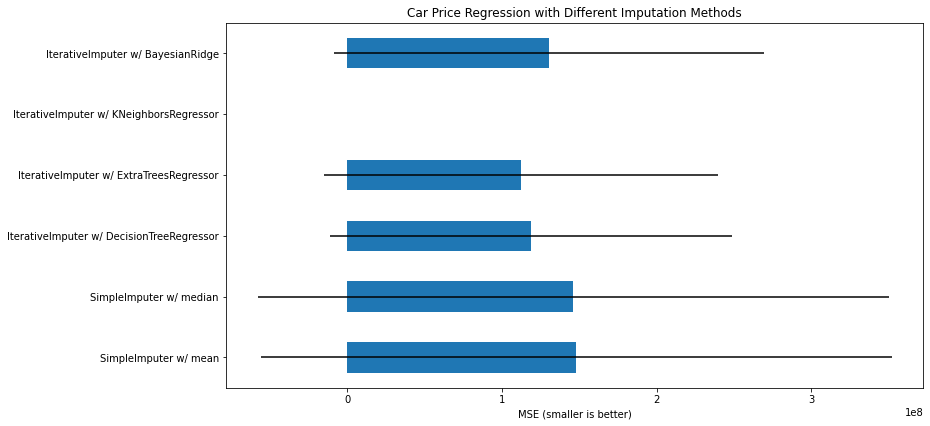
# plot california housing results
fig, ax = plt.subplots(figsize=(13, 6))
means = -scores.mean()
errors = scores.std()
means.plot.barh(xerr=errors, ax=ax)
ax.set_title('Car Price Regression with Different Imputation Methods')
ax.set_xlabel('MSE (smaller is better)')
ax.set_yticks(np.arange(means.shape[0]))
ax.set_yticklabels([" w/ ".join(label) for label in means.index.tolist()])
plt.tight_layout(pad=1)
plt.show()
5. Data Transformation
The following transformation consists of removing the outliers using 2 of the most widely known methods, the Interquartile range and Z Score method. Before applying the Z Score Outlier Removal we firstly need to scale the numeric variables of the data and bring them to have Gaussian(Normal) distribution. Lastly the categorical variables are being encoded.
from sklearn.preprocessing import PowerTransformer
final_pipeline = Pipeline([
('IQR_removal',IQROutlierRemoval(num_att = num_att)),
('std_scaler', StandardScalerIndices()),
('yeo-johnson',PowerTransformerIndices(method='yeo-johnson')),
('z_score_removal', ZScoreOutlierRemoval(num_att = num_att)),
('Imputer', IterativeImputerIndices(estimator = ExtraTreesRegressor(n_estimators=10, random_state=0))),
('join', JoinTransformer(x_train, num_att)), #kategoriski so numericki
('encoder', CategoricalEncoder()),
])
X_train_prepared = pd.DataFrame(final_pipeline.fit_transform(x_train))
joined = X_train_prepared.join(y_train)
y_train = joined['Price']
fig = go.Figure()
fig.add_trace(
go.Histogram(x=X_train_prepared['Mileage'], name='Mileage', visible = True)
)
fig.add_trace(
go.Histogram(x=X_train_prepared['First Registration'], name='First Registration', visible = False)
)
fig.add_trace(
go.Histogram(x=X_train_prepared['Power(hp)'], name='Power(hp)', visible = False)
)
fig.add_trace(
go.Histogram(x=X_train_prepared['Displacement'], name='Displacement', visible = False)
)
fig.update_layout(
updatemenus=[
dict(
active=0,
buttons=list([
dict(label="Mileage",
method="update",
args=[{"visible": [True,False,False,False]},
{"title": "Mileage"
}]),
dict(label="First Registration",
method="update",
args=[{"visible": [False,True,False,False]},
{"title": "First Registration"}]),
dict(label="Power(hp)",
method="update",
args=[{"visible": [False,False,True,False]},
{"title": "Power(hp)"}]),
dict(label="Displacement",
method="update",
args=[{"visible": [False,False,False,True]},
{"title": "Displacement"}]),
]),
)
])
plot(fig, filename = 'fig.html', config = config)
display(HTML('fig.html'))
7. Dimensionality Reduction
7.1 Choosing the right number of components
Next, we are trying different methods of reducing the dimensionality of the data. In order to maximize the accuracy, one of the vital things to keep in mind is to preserve the inital variance of the data. For this, we are plotting the preserved variance against the number of components, for a Principal Component Analyisis(PCA) and Singular Value Decomposition(SVD) model respectively.
pca = decomposition.PCA()
pca.n_components = 121
pca_data = pca.fit_transform(X_train_prepared)
percentage_var_explained = pca.explained_variance_ / np.sum(pca.explained_variance_)
cum_var_explained = np.cumsum(percentage_var_explained)
svd = TruncatedSVD()
svd.n_components = 121
svd_data = svd.fit_transform(X_train_prepared)
percentage_var_explained_svd = svd.explained_variance_ / np.sum(svd.explained_variance_)
cum_var_explained_svd = np.cumsum(percentage_var_explained_svd)
fig = go.Figure()
fig.add_trace(go.Scatter(y = cum_var_explained, name = 'PCA'))
fig.add_trace(go.Scatter(y = cum_var_explained_svd, name = 'SVD'))
fig.update_layout(plot_bgcolor='rgb(255,255,255)',xaxis_title="n_components",
yaxis_title='Variance')
fig.update_xaxes(ticks = 'outside', showline=True, linecolor='black')
fig.update_yaxes(ticks = 'outside', showline=True, linecolor='black')
# Plot figure
plot(fig, filename = 'fig2.html', config = config)
display(HTML('fig2.html'))
7.2 Applying the transformations
Having made the previous analysis we are ready to apply the following dimensionality reduction algorithms:
- Principal Component Analysis(PCA)
- Sparse Principal Component Analysis(SparsePCA)
- Singular Value Decomposition(SVD)
- Isomap
From the plot above it is obvious that we need 86 components to preserve 99% of the inital variance of the data.
pca_pipeline = Pipeline([
('IQR_removal',IQROutlierRemoval(num_att = num_att)),
('std_scaler', StandardScalerIndices()),
('yeo-johnson',PowerTransformerIndices(method='yeo-johnson')),
('z_score_removal', ZScoreOutlierRemoval(num_att = num_att)),
('Imputer', IterativeImputerIndices(estimator = ExtraTreesRegressor(n_estimators=10, random_state=0))),
('join', JoinTransformer(x_train, num_att)), #kategoriski so numericki
('encoder', CategoricalEncoder()),
('pca', decomposition.PCA(n_components = 86))
])
spca_pipeline = Pipeline([
('IQR_removal',IQROutlierRemoval(num_att = num_att)),
('std_scaler', StandardScalerIndices()),
('yeo-johnson',PowerTransformerIndices(method='yeo-johnson')),
('z_score_removal', ZScoreOutlierRemoval(num_att = num_att)),
('Imputer', IterativeImputerIndices(estimator = ExtraTreesRegressor(n_estimators=10, random_state=0))),
('join', JoinTransformer(x_train, num_att)), #kategoriski so numericki
('encoder', CategoricalEncoder()),
('pca', decomposition.SparsePCA())
])
svd_pipeline = Pipeline([
('IQR_removal',IQROutlierRemoval(num_att = num_att)),
('std_scaler', StandardScalerIndices()),
('yeo-johnson',PowerTransformerIndices(method='yeo-johnson')),
('z_score_removal', ZScoreOutlierRemoval(num_att = num_att)),
('Imputer', IterativeImputerIndices(estimator = ExtraTreesRegressor(n_estimators=10, random_state=0))),
('join', JoinTransformer(x_train, num_att)), #kategoriski so numericki
('encoder', CategoricalEncoder()),
('svd', TruncatedSVD(n_components = 86))
])
isomap_pipeline = Pipeline([
('IQR_removal',IQROutlierRemoval(num_att = num_att)),
('std_scaler', StandardScalerIndices()),
('yeo-johnson',PowerTransformerIndices(method='yeo-johnson')),
('z_score_removal', ZScoreOutlierRemoval(num_att = num_att)),
('Imputer', IterativeImputerIndices(estimator = ExtraTreesRegressor(n_estimators=10, random_state=0))),
('join', JoinTransformer(x_train, num_att)), #kategoriski so numericki
('encoder', CategoricalEncoder()),
('isomap', Isomap(n_components = 60))
])
X_train_spca = pd.DataFrame(spca_pipeline.fit_transform(x_train))
X_train_pca = pd.DataFrame(pca_pipeline.fit_transform(x_train))
y_train_pca = y_train.reset_index()
y_train_pca = y_train_pca.drop('index', axis = 1)
X_train_svd = pd.DataFrame(svd_pipeline.fit_transform(x_train))
y_train_svd = y_train_pca.copy()
X_train_isomap = pd.DataFrame(isomap_pipeline.fit_transform(x_train))
y_train_isomap = y_train_pca.copy()
;
8. Additional Feature Engineering
Here, we will try to improve the data quality by grouping the equipment features. This will result in reducing the binary features in our data, thus reducing the data sparcity. The equipment is being classified in the following groups:
- Safety Equipment
- Luxurious Equipment
- General Equipment
- Sensors
- Lights
The same methods of dimensionality reduction are being applied to this newly formed dataset.
safety = ['ABS','Traction control','Driver-side airbag','Side airbag','Passenger-side airbag','Isofix','Immobilizer']
luxury = ['Adaptive Cruise Control','Leather steering wheel','Massage seats','Heated steering wheel','Panorama roof','Touch screen',
'Keyless central door lock','Electrically heated windshield','Alloy wheels','Sunroof','Electrically adjustable seats',
'Navigation system']
general = ['Multi-function steering wheel','Air suspension','Hill Holder','USB','Non-smoking Vehicle','Air conditioning','Automatic climate control','Radio','Bluetooth', 'CD player',
'Power windows','Central door lock','On-board computer','Alarm system', 'Trailer hitch', 'Ski bag','MP3','Digital radio',
'Armrest','Power steering','Electrical side mirrors','Roof rack']
sensors = ['Parking assist system sensors rear','Parking assist system sensors front','Night view assist', 'Blind spot monitor', 'Parking assist system camera'
, 'Parking assist system self-steering','Lane departure warning system','Traffic sign recognition',
'Electronic stability control','Tire pressure monitoring system','Electric tailgate','Rain sensor','Start-stop system']
lights = ['LED Daytime Running Lights','LED Headlights','Adaptive headlights','Daytime running lights','Xenon headlights','Fog lights']
recnik2 = {'luxury':luxury, 'safety':safety, 'general':general, 'sensors':sensors, 'lights':lights}
num_att_new = ['First Registration',
'Mileage',
'Power(hp)',
'Displacement', 'luxury', 'safety', 'general', 'lights', 'sensors']
x_train_red = x_train.copy()
for key in recnik2.keys():
x_train_red
x_train_red[key] = 0
for obj in recnik2[key]:
x_train_red[key] += x_train_red[obj]
x_train_red = x_train_red.drop(obj, axis = 1)
spca_pipeline_reduced = Pipeline([
('IQR_removal',IQROutlierRemoval(num_att = num_att_new)),
('std_scaler', StandardScalerIndices()),
('yeo-johnson',PowerTransformerIndices(method='yeo-johnson')),
('z_score_removal', ZScoreOutlierRemoval(num_att = num_att_new)),
('Imputer', IterativeImputerIndices(estimator = ExtraTreesRegressor(n_estimators=10, random_state=0))),
('join', JoinTransformer(x_train_red, num_att_new)), #kategoriski so numericki
('encoder', CategoricalEncoder()),
#('pca', decomposition.SparsePCA())
])
reduced_x = pd.DataFrame(spca_pipeline_reduced.fit_transform(x_train_red))
joined = reduced_x.join(y_train)
reduced_y = joined['Price']
reduced_reset_y = reduced_y.reset_index()
reduced_reset_y = reduced_reset_y.drop('index', axis = 1 )
;
svd = TruncatedSVD()
svd.n_components = 60
svd_data = svd.fit_transform(reduced_x,reduced_y)
percentage_var_explained_svd = svd.explained_variance_ / np.sum(svd.explained_variance_)
cum_var_explained_svd = np.cumsum(percentage_var_explained_svd)
pca = decomposition.PCA()
pca.n_components = 60
pca_data = pca.fit_transform(reduced_x,reduced_y)
percentage_var_explained = pca.explained_variance_ / np.sum(pca.explained_variance_)
cum_var_explained = np.cumsum(percentage_var_explained)
fig = go.Figure()
fig.add_trace(go.Scatter(y = cum_var_explained, name = 'PCA'))
fig.add_trace(go.Scatter(y = cum_var_explained_svd, name = 'SVD'))
fig.update_layout(plot_bgcolor='rgb(255,255,255)',xaxis_title="n_components",
yaxis_title='Variance')
fig.update_xaxes(ticks = 'outside', showline=True, linecolor='black')
fig.update_yaxes(ticks = 'outside', showline=True, linecolor='black')
# Plot figure
plot(fig, filename = 'fig10.html', config = config)
display(HTML('fig10.html'))
spca_pipeline_reduced2 = Pipeline([
('IQR_removal',IQROutlierRemoval(num_att = num_att_new)),
('std_scaler', StandardScalerIndices()),
('yeo-johnson',PowerTransformerIndices(method='yeo-johnson')),
('z_score_removal', ZScoreOutlierRemoval(num_att = num_att_new)),
('Imputer', IterativeImputerIndices(estimator = ExtraTreesRegressor(n_estimators=10, random_state=0))),
('join', JoinTransformer(x_train_red, num_att_new)), #kategoriski so numericki
('encoder', CategoricalEncoder()),
('pca', decomposition.SparsePCA())
])
pca_pipeline_reduced = Pipeline([
('IQR_removal',IQROutlierRemoval(num_att = num_att_new)),
('std_scaler', StandardScalerIndices()),
('yeo-johnson',PowerTransformerIndices(method='yeo-johnson')),
('z_score_removal', ZScoreOutlierRemoval(num_att = num_att_new)),
('Imputer', IterativeImputerIndices(estimator = ExtraTreesRegressor(n_estimators=10, random_state=0))),
('join', JoinTransformer(x_train_red, num_att_new)), #kategoriski so numericki
('encoder', CategoricalEncoder()),
('pca', decomposition.PCA(n_components = 34))
])
svd_pipeline_reduced = Pipeline([
('IQR_removal',IQROutlierRemoval(num_att = num_att_new)),
('std_scaler', StandardScalerIndices()),
('yeo-johnson',PowerTransformerIndices(method='yeo-johnson')),
('z_score_removal', ZScoreOutlierRemoval(num_att = num_att_new)),
('Imputer', IterativeImputerIndices(estimator = ExtraTreesRegressor(n_estimators=10, random_state=0))),
('join', JoinTransformer(x_train_red, num_att_new)), #kategoriski so numericki
('encoder', CategoricalEncoder()),
('SVD', TruncatedSVD(n_components = 34))
])
reduced_x_spca = pd.DataFrame(spca_pipeline_reduced2.fit_transform(x_train_red))
reduced_x_svd = pd.DataFrame(svd_pipeline_reduced.fit_transform(x_train_red))
reduced_x_pca = pd.DataFrame(pca_pipeline_reduced.fit_transform(x_train_red))
9. Label Encoding instead of One Hot Encoding
When encoding the categorical, independent(input) variables of the data we mainly use the One Hot Encoding techinque where each newly created variable represents one level of the categorical feature. 0 represents absence while 1 represents the presence of that category. We use this approach when handling nominal data, where the categories do not have an inherent order. On the other hand, by using Label Encoder we are imposing ordinality in the data, thus making some levels of the categorical feature more important than others. Unsurprisingly, this might affect some algorithms negatively.
However, when using Tree-Based emseble models this might be just the opposite. These types of algorithms work well with categorical features and there is no difference whether the features are ordinal or nominal.This is because the algorithms does not take the ordinality of the categorical features into account.
final_pipeline = Pipeline([
('IQR_removal',IQROutlierRemoval(num_att = num_att)),
('std_scaler', StandardScalerIndices()),
('yeo-johnson',PowerTransformerIndices(method='yeo-johnson')),
('z_score_removal', ZScoreOutlierRemoval(num_att = num_att)),
('Imputer', IterativeImputerIndices(estimator = ExtraTreesRegressor(n_estimators=10, random_state=0))),
('join', JoinTransformer(x_train, num_att)), #kategoriski so numericki
])
X_train_label = pd.DataFrame(final_pipeline.fit_transform(x_train))
cat = [ 'Make','Model', 'Fuel', 'Body', 'Gearing Type']
encoder = LabelEncoder()
for c in cat:
X_train_label[c] = encoder.fit_transform(X_train_label[c])
spca = decomposition.SparsePCA()
X_train_label_sparse = pd.DataFrame(spca.fit_transform(X_train_label))
grid_no_dim_red = gridSearch(param_grid = {'max_features' : [90,100,110,'auto','sqrt','log2'], 'n_estimators' : [100] },
model = RandomForestRegressor(random_state = 2), X = X_train_prepared, y = y_train)
randomforestCV(max_features = grid_no_dim_red.best_params_['max_features'], n_estimators = grid_no_dim_red.best_params_['n_estimators'],
X = X_train_prepared, y = y_train, message = 'Without dim reduction')
10.1.2 Transformations made:
- Removing outliers with the Interquartile range method
- Standardizing data
- Normalizing the data using the yeo-johnosn power transfromation
- Removing outliers with the Z Score method
- Imputing missing values
- Encoding cateorical variables using One Hot Encoding
- Principal Component Analysis
grid_pca = gridSearch(param_grid = {'n_estimators':[100], 'max_features' : [50,60,70,'auto', 'sqrt', 'log2']},
model = RandomForestRegressor(random_state =2), X = X_train_pca, y = y_train_pca)
randomforestCV(max_features = grid_pca.best_params_['max_features'],
n_estimators = grid_pca.best_params_['n_estimators'],
X = X_train_pca, y = y_train_pca, message = 'PCA applied')
10.1.3 Transformations made:
- Removing outliers with the Interquartile range method
- Standardizing data
- Normalizing the data using the yeo-johnosn power transfromation
- Removing outliers with the Z Score method
- Imputing missing values
- Encoding cateorical variables using One Hot Encoding
- Sparse Principal Component Analysis
grid_spca = gridSearch(param_grid = {'n_estimators':[100], 'max_features' : [50,60,70,80, 'auto','sqrt','log2']},
model = RandomForestRegressor(random_state =2), X = X_train_spca, y = y_train_pca)
randomforestCV(max_features = grid_spca.best_params_['max_features'],
n_estimators = grid_spca.best_params_['n_estimators'],
X = X_train_spca, y = y_train_pca, message = 'SparsePCA applied')
10.1.4 Transformations made:
- Removing outliers with the Interquartile range method
- Standardizing data
- Normalizing the data using the yeo-johnosn power transfromation
- Removing outliers with the Z Score method
- Imputing missing values
- Encoding cateorical variables using One Hot Encoding
- Singular Value Decomposition
grid_svd = gridSearch(param_grid = {'n_estimators':[100], 'max_features' : [40,50,60,70, 'auto','sqrt','log2']},
model = RandomForestRegressor(random_state = 2), X = X_train_svd, y = y_train_svd.values.ravel())
randomforestCV(max_features = grid_svd.best_params_['max_features'],
n_estimators = grid_svd.best_params_['n_estimators'],
X = X_train_svd, y = y_train_svd.values.ravel(), message = 'SVD applied')
10.1.5 Transformations made:
- Grouping the categorical features
- Removing outliers with the Interquartile range method
- Standardizing data
- Normalizing the data using the yeo-johnosn power transfromation
- Removing outliers with the Z Score method
- Imputing missing values
- Encoding cateorical variables using One Hot Encoding
grid_reduced = gridSearch(param_grid = {'n_estimators':[100], 'max_features' : [15,20,25,30,40,50,'auto','sqrt','log2']},
model = RandomForestRegressor(random_state =2), X = reduced_x, y = reduced_y)
randomforestCV(max_features = grid_reduced.best_params_['max_features'],
n_estimators = grid_reduced.best_params_['n_estimators'],
X = reduced_x, y = reduced_y, message = 'Reduced Data Set w/o dim reduction')
10.1.6 Transformations made:
- Grouping the categorical features
- Removing outliers with the Interquartile range method
- Standardizing data
- Normalizing the data using the yeo-johnosn power transfromation
- Removing outliers with the Z Score method
- Imputing missing values
- Encoding cateorical variables using One Hot Encoding
- Principal Component Analysis
grid_reduced_PCA = gridSearch(param_grid = {'n_estimators':[100], 'max_features' : [10,15,20,25,30,'auto','sqrt','log2']},
model = RandomForestRegressor(random_state = 2), X = reduced_x_pca, y = reduced_reset_y.values.ravel())
randomforestCV(max_features = grid_reduced_PCA.best_params_['max_features'],
n_estimators = grid_reduced_PCA.best_params_['n_estimators'],
X = reduced_x_pca, y = reduced_reset_y.values.ravel(), message = 'Reduced Data Set + PCA')
10.1.7 Transformations made:
- Grouping the categorical features
- Removing outliers with the Interquartile range method
- Standardizing data
- Normalizing the data using the yeo-johnosn power transfromation
- Removing outliers with the Z Score method
- Imputing missing values
- Encoding cateorical variables using One Hot Encoding
- Sparse Principal Component Analysis
grid_reduced_SPCA = gridSearch(param_grid = {'n_estimators':[100], 'max_features' : [10,15,20,21,'auto','sqrt','log2',40]},
model = RandomForestRegressor(random_state = 2), X = reduced_x_spca, y = reduced_reset_y.values.ravel())
randomforestCV(max_features = grid_reduced_SPCA.best_params_['max_features'],
n_estimators = grid_reduced_SPCA.best_params_['n_estimators'],
X = reduced_x_spca, y = reduced_reset_y.values.ravel(), message = 'Reduced Data Set + SparsePCA')
10.1.8 Transformations made:
- Grouping the categorical features
- Removing outliers with the Interquartile range method
- Standardizing data
- Normalizing the data using the yeo-johnosn power transfromation
- Removing outliers with the Z Score method
- Imputing missing values
- Encoding cateorical variables using One Hot Encoding
- Singular Value Decomposition
grid_reduced_svd = gridSearch(param_grid = {'n_estimators':[100], 'max_features' : [10,15,20,25,30,'auto','sqrt','log2']},
model = RandomForestRegressor(random_state =42), X = reduced_x_svd, y = reduced_reset_y.values.ravel())
randomforestCV(max_features = grid_reduced_svd.best_params_['max_features'],
n_estimators = grid_reduced_svd.best_params_['n_estimators'],
X = reduced_x_svd, y = reduced_reset_y.values.ravel(), message = 'Reduced Data Set + SVD')
rfr = RandomForestRegressor(random_state = 2,max_depth=17)
rfr.fit(X_train_label,y_train)
rfr_scores = cross_val_score(rfr, X_train_label, y_train, scoring = 'neg_mean_absolute_percentage_error',
cv = 5, n_jobs = -1)
recnik['Method'].append('Label Encoded w/o Dimenisonality Reduction')
recnik['Mean Percentage Error'].append(-rfr_scores.mean())
recnik['Standard Deviation'].append(rfr_scores.std())
xgb = XGBRegressor(random_state = 2)
xgb.fit(X_train_prepared, y_train)
xgb_scores = cross_val_score(xgb, X_train_prepared, y_train, scoring = 'neg_mean_absolute_percentage_error', cv = 5, n_jobs = -1)
print("Scores:", -xgb_scores)
print("Mean:", -xgb_scores.mean())
print("Standard deviation:", xgb_scores.std())
recnik['Method'].append('XGBRegressor w/o dim reduction')
recnik['Mean Percentage Error'].append(-xgb_scores.mean())
recnik['Standard Deviation'].append(xgb_scores.std())
Using the Ensemble Regressor on a data that has been transformed the same way as in part 10.1.9
from sklearn.ensemble import VotingRegressor
vote_mod = VotingRegressor([ ('XGBRegressor', XGBRegressor()),
('RandomForest', RandomForestRegressor(random_state = 2,max_depth=17))])
vote_mod.fit(X_train_label, y_train)
reg_scores = cross_val_score(vote_mod, X_train_label, y_train, scoring = 'neg_mean_absolute_percentage_error',
cv = 5, n_jobs = -1)
print("Scores:", -reg_scores)
print("Mean:", -reg_scores.mean())
print("Standard deviation:", reg_scores.std())
recnik['Method'].append('Voting Regressor w/o dim reduction')
recnik['Mean Percentage Error'].append(-reg_scores.mean())
recnik['Standard Deviation'].append(reg_scores.std())
reg_scores = cross_val_score(vote_mod, X_train_label_sparse, y_train_pca.values.ravel(), scoring = 'neg_mean_absolute_percentage_error',
cv = 5)
print("Scores:", -reg_scores)
print("Mean:", -reg_scores.mean())
print("Standard deviation:", reg_scores.std())
recnik['Method'].append('Voting Regressor + SparsePCA')
recnik['Mean Percentage Error'].append(-reg_scores.mean())
recnik['Standard Deviation'].append(reg_scores.std())
pd.DataFrame.from_dict(recnik)